Useful Scripts¶
Location of the scripts¶
Here are some scripts that you may find useful. They are in the folder “./eppy/useful_scripts”
And now for some housekeeping before we start off
[1]:
import os
os.chdir("../eppy/useful_scripts")
# changes directory, so we are where the scripts are located
[2]:
# you would normaly install eppy by doing
# python setup.py install
# or
# pip install eppy
# or
# easy_install eppy
# if you have not done so, the following three lines are needed
import sys
# pathnameto_eppy = 'c:/eppy'
pathnameto_eppy = '../../'
sys.path.append(pathnameto_eppy)
If you look in the folder “./eppy/useful_scripts”, you fill find the following scripts
The scripts are:
- eppy_version.py
- idfdiff.py
- loopdiagram.py
- eppyreadtest_folder.py
- eppyreadtest_file.py
eppy_version.py¶
Many scripts will print out some help information, if you use the –help option. Let us try that
[3]:
%%bash
# ignore the line above. It simply lets me run a command line from ipython notebook
python eppy_version.py --help
usage: eppy_version.py [-h]
I print the current version of eppy. Being polite, I also say hello !
optional arguments:
-h, --help show this help message and exit
That was useful !
Now let us try running the program
[4]:
%%bash
# ignore the line above. It simply lets me run a command line from ipython notebook
python eppy_version.py
Hello! I am eppy version 0.5.52
Redirecting output to a file¶
Most scripts will print the output to a terminal. Sometimes we want to send the output to a file, so that we can save it for posterity. We can do that py using “>” with the filename after that. For eppy_version.py, it will look like this:
python eppy_version.py > save_output.txtSome of the following scripts will generate csv or html outputs. We can direct the output to a file with .html extension and open it in a browser
Compare two idf files - idfdiff.py¶
This script will compare two idf files. The results will be displayed printed in “csv” format or in “html” format.
You would run the script from the command line. This would be the terminal on Mac or unix, and the dos prompt on windows. Let us look at the help for this script, by typing:
[8]:
%%bash
# ignore the line above. It simply lets me run a command line from ipython notebook
python idfdiff.py -h
usage: idfdiff.py [-h] [--idd IDD] (--csv | --html) file1 file2
Do a diff between two idf files. Prints the diff in csv or html file format.
You can redirect the output to a file and open the file using as a spreadsheet
or by using a browser
positional arguments:
file1 location of first with idf files = ./somewhere/f1.idf
file2 location of second with idf files = ./somewhere/f2.idf
optional arguments:
-h, --help show this help message and exit
--idd IDD location of idd file = ./somewhere/eplusv8-0-1.idd
--csv
--html
Now let us try this with two “idf” files that are slightly different. If we open them in a file comparing software, it would look like this:
[9]:
from eppy.useful_scripts import doc_images #no need to know this code, it just shows the image below
for_images = doc_images
for_images.display_png(for_images.filemerge) # display the image below
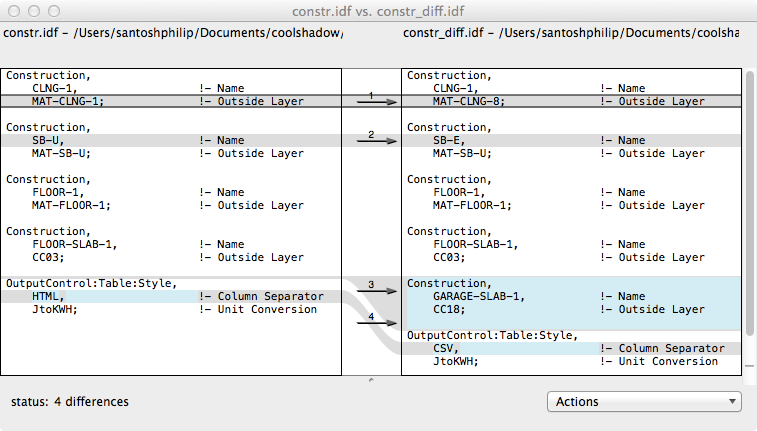
There are 4 differences between the files. Let us see what idfdiff.py does with the two files. We will use the –html option to print out the diff in html format.
[11]:
%%bash
# python idfdiff.py idd file1 file2
python idfdiff.py --html --idd ../resources/iddfiles/Energy+V7_2_0.idd ../resources/idffiles/V_7_2/constructions.idf ../resources/idffiles/V_7_2/constructions_diff.idf
../resources/iddfiles/Energy+V7_2_0.idd ../resources/idffiles/V_7_2/constructions.idf ../resources/idffiles/V_7_2/constructions_diff.idf
<html><p>file1 = ../resources/idffiles/V_7_2/constructions.idf</p><p>file2 = ../resources/idffiles/V_7_2/constructions_diff.idf</p><table border="1"><tr><th>Object Key</th><th> Object Name</th><th> Field Name</th><th> file1</th><th> file2</th></tr><tr><td>MATERIAL</td><td>F08 Metal surface</td><td></td><td>is here</td><td>not here</td></tr><tr><td>MATERIAL</td><td>F08 Metal surface haha</td><td></td><td>not here</td><td>is here</td></tr><tr><td>MATERIAL</td><td>G05 25mm wood</td><td>Conductivity</td><td>0.15</td><td>0.155</td></tr><tr><td>CONSTRUCTION</td><td>Exterior Door</td><td>Outside Layer</td><td>F08 Metal surface</td><td>F08 Metal surface haha</td></tr></table></html>
idfdiff.py:193: UserWarning: No parser was explicitly specified, so I'm using the best available HTML parser for this system ("lxml"). This usually isn't a problem, but if you run this code on another system, or in a different virtual environment, it may use a different parser and behave differently.
The code that caused this warning is on line 193 of the file idfdiff.py. To get rid of this warning, pass the additional argument 'features="lxml"' to the BeautifulSoup constructor.
soup = BeautifulSoup()
file1 = ../resources/idffiles/V_7_2/constructions.idf
file2 = ../resources/idffiles/V_7_2/constructions_diff.idf
| Object Key | Object Name | Field Name | file1 | file2 |
|---|---|---|---|---|
| MATERIAL | F08 Metal surface | not here | is here | |
| MATERIAL | F08 Metal surface haha | is here | not here | |
| MATERIAL | G05 25mm wood | Conductivity | 0.15 | 0.155 |
| CONSTRUCTION | Exterior Door | Outside Layer | F08 Metal surface | F08 Metal surface haha |
It does look like html :-). We need to redirect this output to a file and then open the file in a browser to see what it looks like. Displayed below is the html file
[12]:
from eppy.useful_scripts import doc_images #no need to know this code, it just shows the image below
from IPython.display import HTML
h = HTML(open(doc_images.idfdiff_path, 'r').read())
h
[12]:
file1 = ../resources/idffiles/V_7_2/constr.idf
file2 = ../resources/idffiles/V_7_2/constr_diff.idf
| Object Key | Object Name | Field Name | file1 | file2 |
|---|---|---|---|---|
| CONSTRUCTION | CLNG-1 | Outside Layer | MAT-CLNG-1 | MAT-CLNG-8 |
| CONSTRUCTION | GARAGE-SLAB-1 | is here | not here | |
| CONSTRUCTION | SB-E | is here | not here | |
| CONSTRUCTION | SB-U | not here | is here | |
| OUTPUTCONTROL:TABLE:STYLE | Column Separator | HTML | CSV |
Pretty straight forward. Scroll up and look at the origin text files, and see how idfdiff.py understands the difference
Now let us try the same thin in csv format
[14]:
%%bash
# python idfdiff.py idd file1 file2
python idfdiff.py --csv --idd ../resources/iddfiles/Energy+V7_2_0.idd ../resources/idffiles/V_7_2/constr.idf ../resources/idffiles/V_7_2/constr_diff.idf
../resources/iddfiles/Energy+V7_2_0.idd ../resources/idffiles/V_7_2/constr.idf ../resources/idffiles/V_7_2/constr_diff.idf
file1 = ../resources/idffiles/V_7_2/constr.idf
file2 = ../resources/idffiles/V_7_2/constr_diff.idf
Object Key, Object Name, Field Name, file1, file2
CONSTRUCTION,CLNG-1,Outside Layer,MAT-CLNG-1,MAT-CLNG-8
CONSTRUCTION,GARAGE-SLAB-1,,not here,is here
CONSTRUCTION,SB-E,,not here,is here
CONSTRUCTION,SB-U,,is here,not here
OUTPUTCONTROL:TABLE:STYLE, ,Column Separator,HTML,CSV
We see the same output, but now in csv format. You can redirect it to a “.csv” file and open it up as a spreadsheet
loopdiagram.py¶
Diagrams of HVAC loops¶
This script will draw all the loops in an idf file. It is a bit of a hack. So it will work on most files, but sometimes it will not :-(. But it is pretty useful when it works.
If it does not work, send us the idf file and we’ll try to fix the code
Make sure grapphviz is installed for this script to work
Again, we’ll have to run the script from the terminal. Let us look at the help for this script
[15]:
%%bash
# ignore the line above. It simply lets me run a command line from ipython notebook
python loopdiagram.py --help
Couldn't import dot_parser, loading of dot files will not be possible.
usage: loopdiagram.py [-h] idd file
Draw all the loops in the IDF file.
There are two output files saved in the same location as the idf file:
- idf_file_location/idf_filename.dot
- idf_file_location/idf_filename.png
positional arguments:
idd location of idd file = ./somewhere/eplusv8-0-1.idd
file location of idf file = ./somewhere/f1.idf
optional arguments:
-h, --help show this help message and exit
Pretty straightforward. Simply open png file and you will see the loop diagram. (ignore the dot file for now. it will be documented later)
So let us try this out with and simple example file. We have a very simple plant loop in “../resources/idffiles/V_7_2/plantloop.idf”
[16]:
%%bash
# ignore the line above. It simply lets me run a command line from ipython notebook
python loopdiagram.py ../resources/iddfiles/Energy+V7_2_0.idd ../resources/idffiles/V_7_2/plantloop.idf
Couldn't import dot_parser, loading of dot files will not be possible.
constructing the loops
cleaning edges
making the diagram
saved file: ../resources/idffiles/V_7_2/plantloop.dot
saved file: ../resources/idffiles/V_7_2/plantloop.png
The script prints out it’s progress. On larger files, this might take a few seconds. If we open this file, it will look like the diagram below
Note: the supply and demnd sides are not connected in the diagram, but shown seperately for clarity
[17]:
from eppy.useful_scripts import doc_images #no need to know this code, it just shows the image below
for_images = doc_images
for_images.display_png(for_images.plantloop) # display the image below
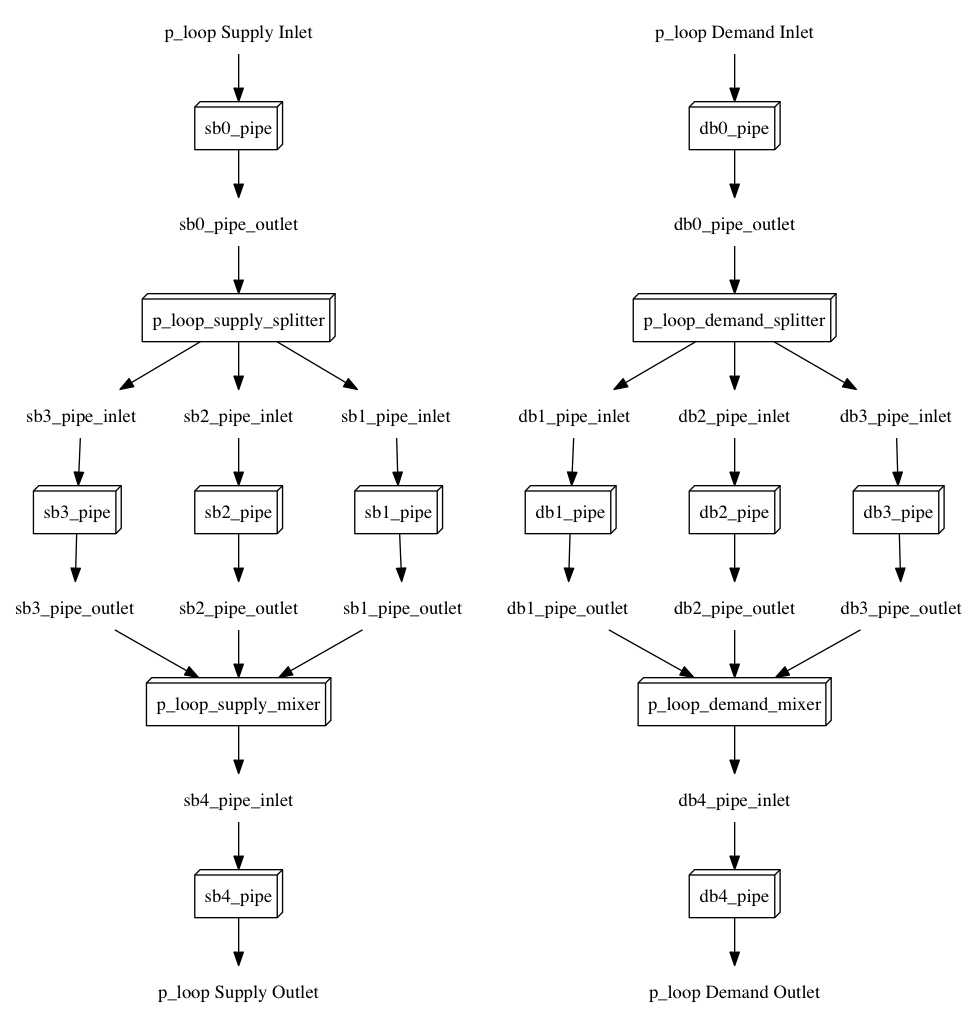
That diagram is not a real system. Does this script really work ?
Try it yourself. Draw the daigram for “../resources/idffiles/V_7_2/5ZoneCAVtoVAVWarmestTempFlow.idf”
Names in loopdiagrams¶
Designbuilder is an energyplus editor autogenerates object names like “MyHouse:SAPZone1”
Note the “:” in the name.
Unfortunatley “:” is a reserved character when making a loop diagrams. (eppy uses pydot and grapphviz which has this constraint)
to work around this, loopdiagram will replace all “:” with a “__”
So the names in the diagram will not match the names in your file, but you can make out what is going on
eppyreadtest_folder.py¶
Not yet documented
eppyreadtest_file.py¶
Not yet documented